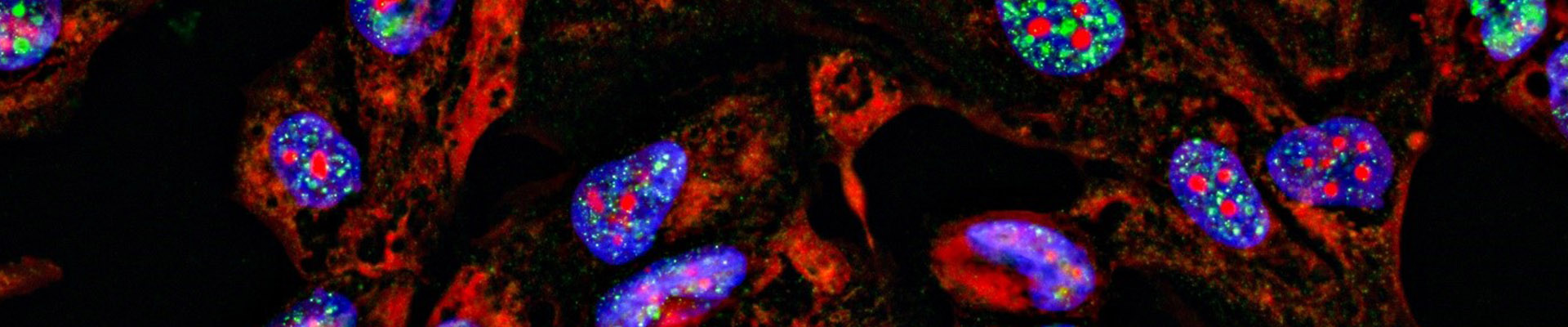
Protein Subcellular Localization
Online InquiryAs a good partner of pharmaceutical companies and research institutions, Creative Proteomics has rich experience in subcellular localization. Subcellular localization refers to the specific location of a protein or a gene expression product in a cell. Only when proteins are transported to specific organelles after being guided by the sorting signal can they participate in various life activities of the cell and perform its functions. If the transport location is wrong, it will affect the cell function and even the whole organism. Therefore, the correct localization of proteins in cells is the guarantee of the orderly operation of cellular systems. Creative Proteomics provides subcellular localization prediction and identification services for the study of protein structure, properties, function, protein-protein interactions, disease mechanisms, and the development of new drugs.
Advantages of Protein Subcellular Localization:
- Rich proteomics experience. The company has been engaged in proteomics related services for 16 years.
- According to different samples and requirements, we provide a variety of subcellular localization methods to choose from.
- Custom services. Based on your samples and requirements, we can customize a dedicated solution for you.
- Efficient, fast and cost-effective.
Methods of Protein Subcellular Localization Services We Can Provide:
We provide a variety of techniques for analyzing subcellular localization, you can choose according to your sample and needs.
1. Immunofluorescent Labeling Service
Immunofluorescence has applications in a variety of ways, including detecting the expression of endogenous proteins in cells, observing the subcellular localization of proteins, and co-localizing the two proteins. And the precise subcellular localization of endogenous and exogenous over-expressed proteins (including cell membranes, mitochondria, lysosomes, nuclear membranes, nucleoli and other organelles). Direct immunofluorescence and indirect immunofluorescence are available. This method reflects the location of the target protein in the cell in a living state, which is more direct and accurate. However, this method requires antibodies specific to the target protein.
2. Fluorescent Protein Fusion Service
This method has strong applicability and short cycle. It can be applied to real-time positioning and dynamic research of living bodies, with high sensitivity and high throughput. Localization results may differ from the protein distribution under in vivo conditions. The over- or under-expression of the fusion protein may make the localization results unclear, especially the localization of membrane proteins such as the plasma membrane. From vector construction to protein subcellular localization, Creative Proteomics provides a one-stop service.
3. Subcellular Localization Prediction Service
Large-scale high-throughput sequencing technology has led to an exponential increase in the number of protein sequences. Facing these huge amounts of data, advanced and efficient computer automated data processing technology is needed. Our computer predictions provide the following three algorithms: neural networks, support vector machines, and nearest neighbor algorithms. To ensure that the results are more rigorous and reliable, we will use leaving a cross-validation to check the accuracy of the algorithm.
* For Research Use Only. Not for use in diagnostic procedures.