Exosome Marker- Introduction and Identification
Online InquiryWhat Are Exosome Markers?
Exosomes are extracellular vesicles produced by all cells and are about 40-120 nm in size (average about 100 nm). Almost all types of cells can secrete exosomes, and also exosomes are widely found in body fluids, including blood, tears, urine, saliva, breast milk, and ascites.
Exosomes are rich in nucleic acids (microRNA, lncRNA, circRNA, mRNA, tRNA, etc.), proteins, lipids, and metabolites, and serve as a medium for intercellular communication. Exosomes have important functions in many physiological and pathological processes such as immune surveillance, inflammatory response and cancer development, especially in intercellular communication, recoding of genes in target cells, and tumor development and invasive metastasis.
The proteins commonly enriched in exosomes are Rab GTPases, SNAREs, flotillin, Alix, TSG101 and tetraspanins. Meanwhile, CD9, CD63 and CD81 are abundant in exosomes and are widely considered as markers of exosomes.
Exosomes can secrete a variety of membrane and cytoplasmic proteins regardless of the normal or pathological state of the organism. Therefore, exosomal proteins may be used for clinical diagnosis.
Function of Exosome Protein Marker
CD9, CD63, and CD81 are all members of the TM4SF/ Tetraspanins family of membrane proteins with a quadruple transmembrane structure, capable of aggregating on the surface of cell membranes and often used as surface markers for exosomes.
CD9
CD9 is expressed in a variety of hematopoietic and non-hematopoietic cells, such as stromal cells, megakaryocytes, platelets, B and T lymphocytes, dendritic cells, endothelial cells, mast cells, eosinophils, and basophils. Localized to extracellular or extracellular bodies, cell membranes.
In macrophages, CD9 functions in association with CD81 and β-1 and β-2 integrins and prevents macrophage fusion into multinucleated giant cells.CD9 is an integrin-associated integral membrane protein that regulates different processes such as sperm-egg fusion, platelet activation and aggregation, and cell adhesion. CD9 is now also considered as an anti-inflammatory marker for monocytes and macrophages.
CD63
CD63 can be detected in platelets (at the protein level), as well as in dysplastic nevi, radial growth-phase primary melanomas, hematopoietic cells, and tissue macrophages. It is localized in cell membranes, intracellular bodies, and lysosomes.
CD63 regulates extracellular vesicle production and sorting of intracellular body cargo. cd63+ exosomes are significantly increased in patients with melanoma and other cancers. Therefore, CD63 has been suggested as a protein marker for cancer.
CD81
CD81 is expressed in B cells, monocytes, macrophages, initial CD4+ T cells, and memory CD4+ T cells. Localized to the cell membrane, basement membrane.
CD81 can form complexes with other four transmembrane proteins, integrins, co-receptors, and MHC class I or II molecules to affect B and T cell adhesion, morphology, activation, proliferation, and differentiation. CD81 was shown to be significantly increased in the sera of patients with chronic hepatitis C, suggesting that CD81 can be used as a diagnostic marker for hepatitis C virus infection.
How Do You Identify Exosomes?
There are 3 markers for exosome identification: transmission electron microscope (TEM), nanoparticle tracking analysis (NTA), and protein markers.
The current traditional methods for analyzing exosomal proteins are western blot, mass spectrometry, scanning electron microscopy, and flow analysis.
Western Blot is most commonly used to experimentally identify exosomal characterization protein markers. The experimental procedure includes exosome protein extraction, marker selection, western blot experiment and result analysis.
Exosome proteins are extracted and exosome protein concentrations are determined by the BCA method. Then WB experiment is performed. Combined with the literature, suitable markers were found and positive and negative controls were set. Commonly used markers include CD9, CD63, CD81, Alix, Tsg101, Calnexin. calnexin is a negative control, which is present in whole cells but not in exosomes; the others are positive controls, which are present in both whole cells and exosomes. In addition, when selecting positive indicators for exosomes, 2-3 indicators are generally recommended for validation to ensure the accuracy of the results.
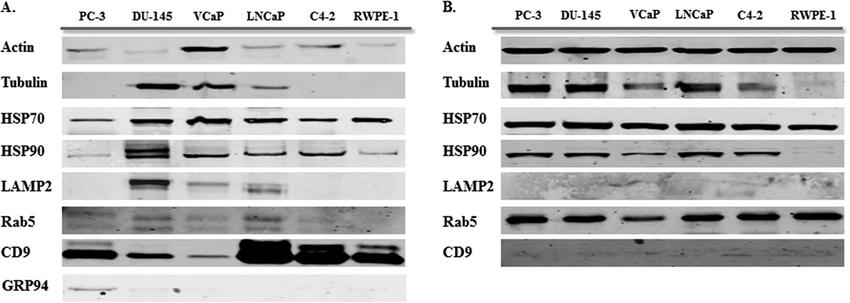 Western blot analysis for exosome markers in exosomes and corresponding cell lysate samples (Hosseini et al., 2012).
Western blot analysis for exosome markers in exosomes and corresponding cell lysate samples (Hosseini et al., 2012).
For a more precise identification of exosomes, the location of proteins in exosomes needs to be observed by exosome immunocolloidal gold electron microscopy. Modern advanced scanning electron microscopy has reached a resolution of about 1 nm, which is sufficient for exosome size measurement. In exosome studies, it is possible to directly observe the morphology of exosomes in the sample.
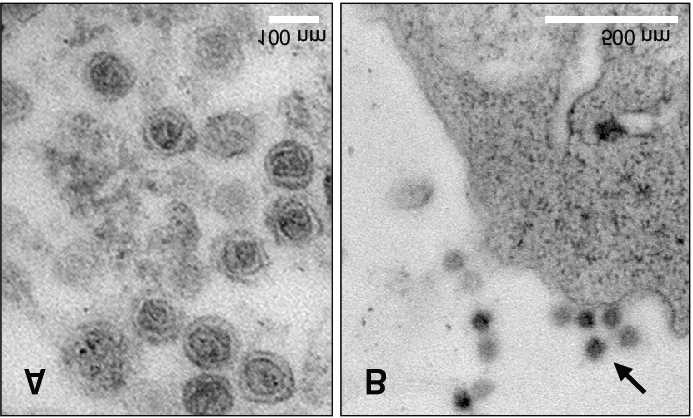 Cryotransmission electron microscopy (TEM) image (Choi et al., 2017)
Cryotransmission electron microscopy (TEM) image (Choi et al., 2017)
NTA ensures the original state of exosomes, fast detection, and provides information on exosome particle size and concentration after detection.
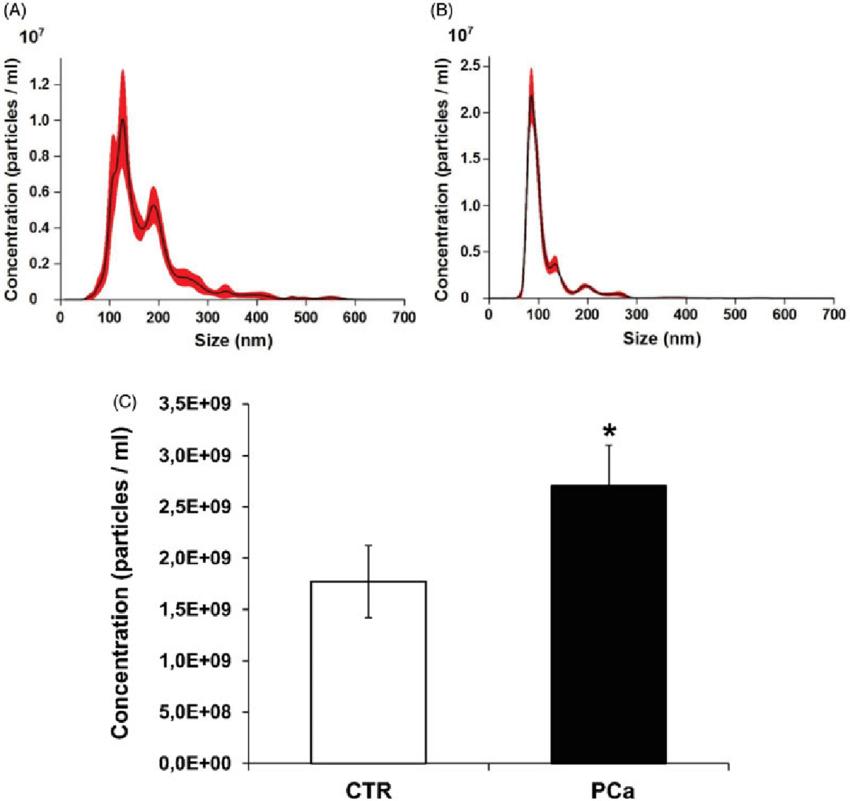 Nanoparticle tracking analysis (NTA) quantification of exosomes (Logozzi et al., 2020)
Nanoparticle tracking analysis (NTA) quantification of exosomes (Logozzi et al., 2020)
Creative Proteomics offers a fast and efficient one-stop service for exosome extraction and purification as well as exosome protein marker analysis. All you need to do is provide samples and we will deliver a report of your satisfactory results.
References
- Hosseini-Beheshti, et al. (2012). Exosomes as biomarker enriched microvesicles: characterization of exosomal proteins derived from a panel of prostate cell lines with distinct AR phenotypes. Molecular & Cellular Proteomics, 11(10), 863-885.
- Choi, H., & Mun, J. Y. (2017). Structural analysis of exosomes using different types of electron microscopy. Applied Microscopy, 47(3), 171-175.
- Logozzi, M., et al. (2020). Plasmatic exosomes from prostate cancer patients show increased carbonic anhydrase IX expression and activity and low pH. Journal of Enzyme Inhibition and Medicinal Chemistry, 35(1), 280-288.
* For Research Use Only. Not for use in diagnostic procedures.